How to evolve graphs with Genetic
Programming
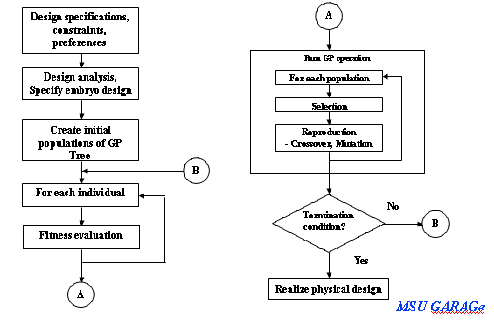
Introduction Concepts Procedure
Evolving graphs by evolutionary algorithms can be classified into two
classes:
one is direct encoding, specifying the
connection and node information directly in the genome with a specific data
structure. Another is indirect encoding or generative encoding, which
encodes a graph growing/development program as the genotype, the phenotype
graphs are generated by executing the developmental programs. We are interested
in the second approach, namely developmental genetic
programming approach for graph evolving, since all significant
achievements of genetic programming by professor John. Koza is made in this way.
And a generative encoding is a much more biologically feasible strategy.
Basic concepts for graph evolving
Embryo graph: A starting graph where later topological manipulation in the GP program can work on. A set of Modifiable sites need to be defined to show places that GP program can modify.

GP function set for graph evolving: the set of function building blocks for genetic programming. Here, it could be the wrappers for topology manipulations, it could be logical operations (and, or), or arithmetic functions or constructs that exist in a programming language.
|
Here are some operators for graph operation in EvoGraph
|
Here are some GP topology operators in
Bondgraph Evolution
|
Modifiable Sites (or Writehead): A set of marks in the structures, indicating where the topology manipulating GP functions can work to modify and transform an embro graph into a design solution.

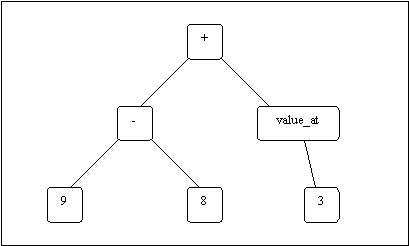
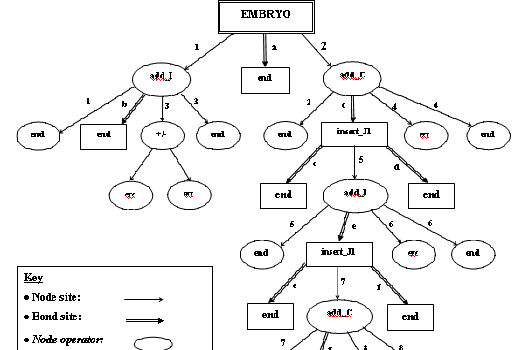
GP tree: A parse tree type struct composed of GP function and termimal buildig blocks, whose execution will lead to the development of a phenotype. GP tree can be modified by crossover and mutation, which will indirectly modify the topology as the phenotype derived from GP tree execution.
| A simple arithmatic GP tree | A somewhat complex bondgraph constructing tree |
 |
 |
GP individual: .Usually a set of GP trees, whose evaluation will lead to a full phenotype to which performance fitness can be evaluated. In most cases, only a single tree is used for a GP individual
Strongly Typed GP: genetic programming with type information attached to each GP terminals and functions, so that in GP crossover and mutation, syntactically correct trees can be ensured.
for example, EvoGraph uses a strongly typed lilGP to do topology synthesis, since nodes and edges ask for different set of topology operations, they have to be typed as edge opertors, node operators, arithmetic operators, etc.